GENÓMICA DE POBLACIONES HUMANAS Y ENFERMEDADES
|
“Human genetics, as a field of genetics that lacks the benefit of an experimental system, therefore has a special reliance on population genetic theory”.
|
- Responsable: Profesor Hernán Dopazo
- Colaboradores: Lic. Juan Manuel Berros, Lic. Nicolás Moreyra, Dr. Diego de Panis y Dr. Jeremias Zubrzycki.
- Duración: 90 hs. Teóricas y Prácticas
THEORY |
|
Theme 1: Personal genomes & NGS Technologies
The Human Genome Project (Public and Private). Diploid Genomes: Venter, & Watson. Comparing Genomes: Europeans, Asians, and Africans. NGS Technologies: Roche/454, Illumina, Helicos, Pacific Bioscience, Ion Torrent/Proton and Oxford Nanopore. Paired-end & Mate-Pair libraries. Multiplex sequencing. Genomic Enrichment Strategies. The Big-Data Storage Problem.
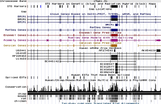
Theme 2: Human genome variability
On Human Variation. Single Nucleotide Polymorphisms (SNPs). Tag-SNPs. Haplotypes and Linkage Disequilibrium in the Human Genome. Haplotype Blocks. Recombination Hotspots. The HapMap Project. Phase I, II & III. Genotype imputation. The 1,000 Genomes Project: Pilot, Phase II and III. The Encode Project Phase I and II.
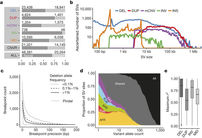
Theme 3: STRUCtural variants & populations
Structural Variants (SVs) in the Human Genome. Genotyping and NGS Methods. Mapping Strategies of NGS-SVs. CNVs formation mechanisms. The SVs Databases. SVs in Human Populations. Population Genetics of SVs. The Human Pan-Genome Concept.
Theme 4: Human demography & population Structure
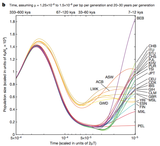
Shared & Private population variants. Deep sequencing projects. Demographic Explosion of Human Populations. Models of Modern Human Origins. Genomic Evidences of the Out-of-Africa Serial Founder Effect Model. Population Genetic Structure of Africans, Europeans and Americans. Human Races: What they mean?
Theme 5: Test of neutrality & Signatures of selection
Tajima’s D; Fay & Wu’s H; MK; dN/dS and LRH test. Ancient and Recent Signatures of Selection in the Human Genome. Archaic Adaptive Introgression in Humans. Natural Selection on Functional Modules.
Theme 6: Genomics & Human DISEASES
Genetic Diseases. Relative Risk. Odds-Ratio. Parametric Linkage Analysis. LOD Score. Sensitivity & Specificity. Population & Family Association Studies. Population Stratification. The Genetic Architecture of Human Diseases. Loss-of-Function (LoF) Variants. The Distribution of the Fitness Effects. The Human Disease Network and the Interactome.
Theme 7: Genome-WIDE Association studies
Heritability. Common Disease / Common Variants Hypothesis & Alternative Hypothesis. GWAS. Replicability. The Enigma of the Hidden Heritability. The Genetic Architecture of Quantitative Characters. The Human & Drosophila Comparison.
Theme 8: PERSONALISED & Precision medicine
The Molecular Medical Diagnostic (MDx) Landscape. Direct to Consumer (DTC) Testing. Heterozygous Carries of Mendelian Diseases. Pre-implantational Genetic Screening (PGS) & Diagnostic (PGD). Karyomapping. Prenatal & Postnatal Genomic Tests. Gene Panels. Exome Sequencing. Non-Invasive Genomic Tests. Single Cell Genomics. CRISPR-Cas9 System & Genome Edition.
HANDs on computers Excercises |
|
TP1- Exploring Human Genome Data
Genomic databases. Ensembl structure and use for variants search. Using NCBI MapViewer browser. Access and data retrieve using BioMart, and the basic uses of the Perl Application Program Interfaces (API).
TP2- INtroduction to Linux using ubuntu
Overview of the file system. Users, groups and permissions. The command-line interface. Basic commands and scripting. Running Local Blast example.
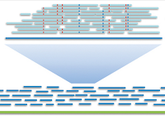
TP3- NGS methods & Quality control
Introduction & Technical Evolution of Next Generation Sequencing. Quality control of NGS reads using FastQC. Processing of reads using FASTX-toolkit & cutadapt.
TP4- Genome AssemblING WITH NGS DATA
Strategies and methods. De novo assembly and evaluation. Reference’s data alignment and visualization.
TP5- INTRODUCTION TO RNAseq
Strategies and methods. Transcriptome assembly. Mapping and counting. Differential expression analysis.
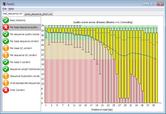
TP6- POPULATION GENOMICS USING 1KGP DATA
Exploring the 1KG browser. Visualization and download. Linkage disequilibrium, Tagging SNP’s and Haploview. Variant Call File (VCF) format. Download and analysis of 1000G data with tabix and VCF tools.
TP7- HUMAN POPULATION STRUCTURE
Filter SNPs in LD. Explore HWE predictions. Computing and plotting MAF frequency spectra. Estimation of ancestries using ADMIXTURE. Principal components analysis. Phenotype association and Manhattan plots.
TP8- SIGNATURES OF SELECTION
Find Signatures of Selection in the Human Genome using 1000 Genomes Selection Browser, Fst, PBS and iHS statistics.
TP9- GWAS
GWAS using Plink 1.9
TP10- VARIANT CALLING AND ANNOTATION
GATK Best Practices Pipeline: data pre proccesing, local realignment around indels, base quality score recalibration, variant discovery.